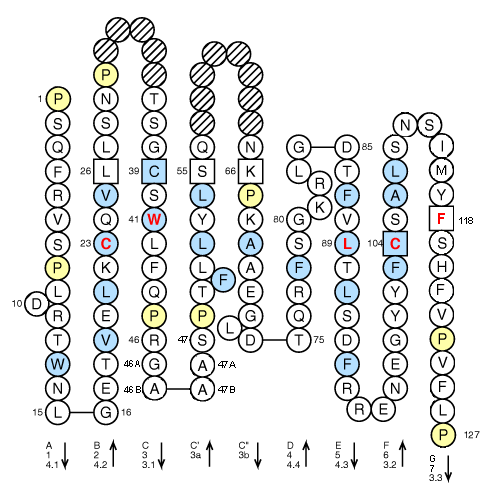
The candidate conserved residues with the highest S cr is selected, and the CDR3 region is located between the two conserved residues in V and J gene segments. Denoting the distance between a matching 3-string to the conserved residue by d k, the support score is calculated as: Σ(1/d k). 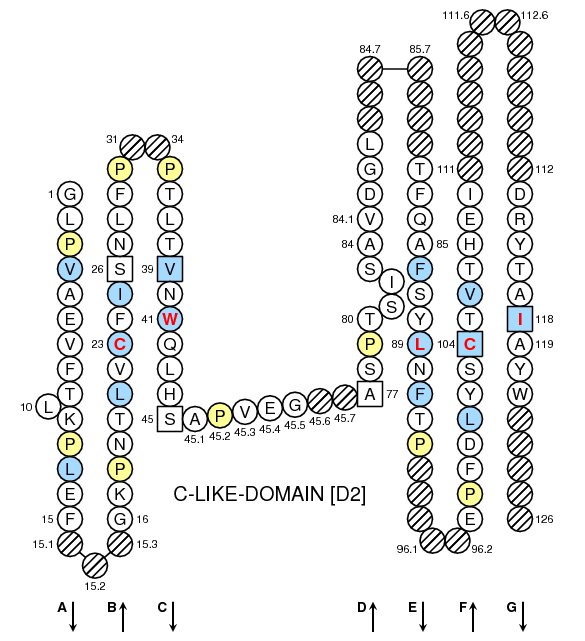
To quantify the concept of “being supported by most 3-strings,” we defined a numeric metric named “conserved residue support score (S cr)” for each residue on query sequence. To avoid the disturbance caused by repeated 3-strings, the position of the conserved residue was determined by the number of supported 3-strings. Then the position of conserved residue (“C” for V segment and “F(GXG) or W(GXG)” for J segment) on the query sequence is determined by the position of the matched 3-strings. The translation frame and the reference profiles with the most N mk were selected for following analysis. First, query sequences are translated on all six frames and compared with the 3-string profiles of reference V and J amino acids sequences. The algorithm of CDR3 identification has two stages. The whole process misses the V and J sequences without CDR3, which may cause bias for V and J annotations because using only the part of the V and J sequence contained within CDR3 is not adequate for annotate-specific V/J segments. To find the CDR3 sequence, the algorithm of MiTCR separates high- and low-quality reads in the first step, and then merges low-quality reads to high-quality CDR3 clusters. The MiTCR is a recently released software that can process massive sequencing reads ( 23). It is implemented using Python language that is much slower than the compiled languages, and its accuracy is only 88%, which is not optimal. The Decombinator is a toolkit for analyzing TCR from short reads ( 22). It is based on pyromap that is specifically designed for 454 pyrosequencing platform ( 21) thus, it does not fit with other sequencing platforms. In contrast, the IRmap is a program that maps the sequencing reads to reference V and J gene segments ( 5). However, the program can only analyze the assembled paired-end reads, and no software was released to the public. The algorithm is based on sequencing reads alignment with reference V gene and J gene segments for the identification of TCR sequences, as well as applying a 96% cutoff value on J gene segment diversity. More recently, the iSSAKE-like strategy was developed by Warren et al. Because of the sequencing cost, short paired-end reads generated by next-generation sequencing technology are and will remain the main source of data, and we believe that the TCRklass is a useful and reliable toolkit for TCR repertoire analysis. We applied TCRklass on large datasets of two human and three mouse TCR repertoires it demonstrated higher reliability on CDR3 identification and much less biased V/J profiling, which are the two components contributing the diversity of the repertoire. We tested TCRklass using manually curated short read datasets in comparison with in silico datasets it showed higher precision and recall rates on CDR3 identification. To decipher the complexity of TCR repertoire, we developed an integrated pipeline, TCRklass, using K-string–based algorithm that has significantly improved the accuracy and performance over existing tools. The next-generation sequencing technology has promoted the study on human TCR repertoire, which is essential for the adaptive immunity.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
